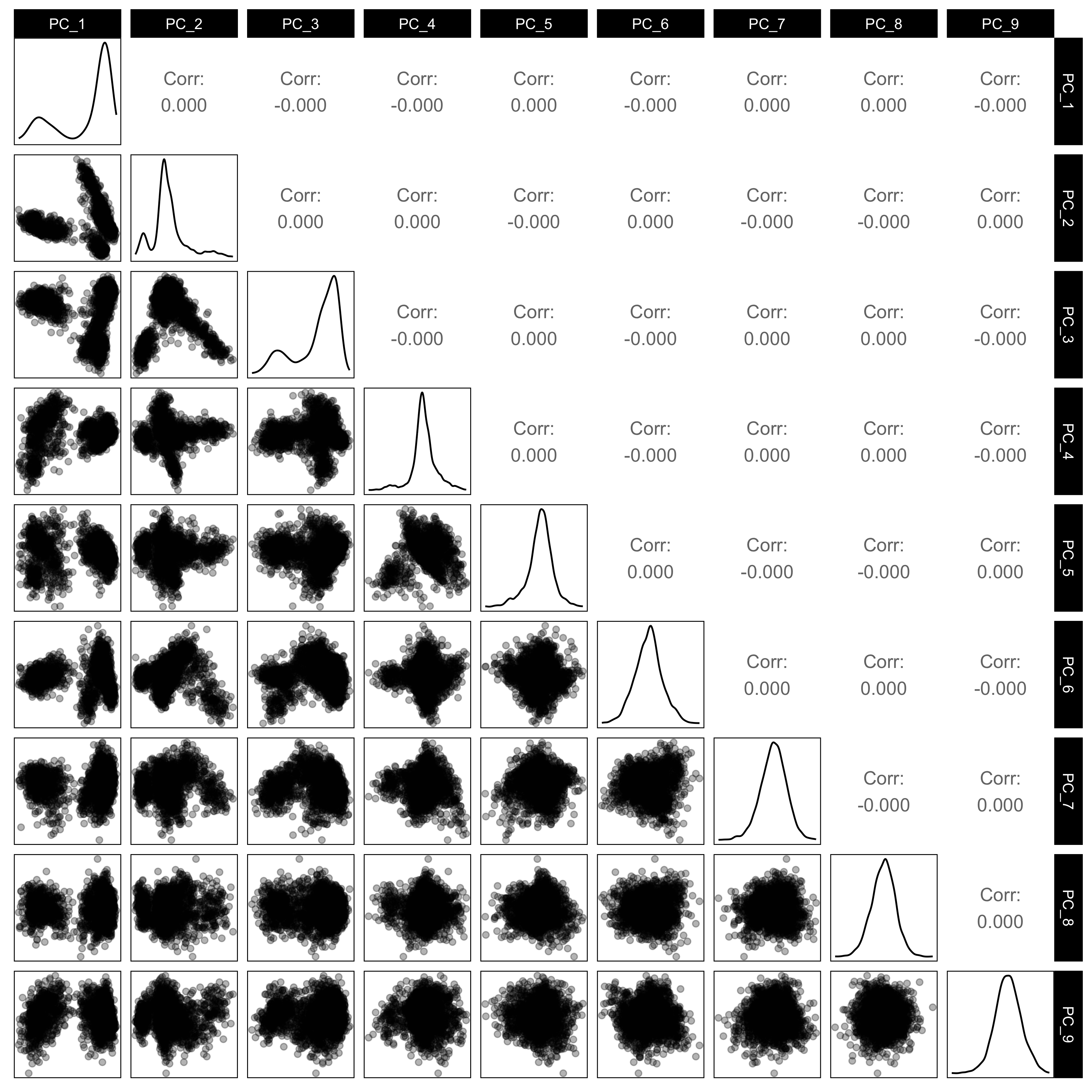
Visualising your fitted non-linear dimension reduction model in the high-dimensional data space
Joint work with Prof. Dianne Cook, Dr. Paul Harrison, Dr. Michael Lydeamore, Dr. Thiyanga S. Talagala
High-dimensional data
\(X = \begin{bmatrix} \textbf{x}_{11} & \textbf{x}_{12} & \cdots & \textbf{x}_{1p}\\ \textbf{x}_{21} & \textbf{x}_{22} & \cdots & \textbf{x}_{2p} \\\vdots & \vdots & \ddots & \vdots & \\ \textbf{x}_{n1} & \textbf{x}_{n2} & \cdots & \textbf{x}_{np}\\\end{bmatrix}\)
Peripheral Blood Mononuclear Cells (PBMC)
Tour
What is a tour?
- Interactive and dynamic graphics to visualise high-dimensional data.
Why is the tour technique employed?
Tour shows a sequence of linear projections as a movie.
It involves mentally assembling multiple low-dimensional views to comprehend the structure in higher dimensions.
Software: langevitour
Non-linear dimensional reduction (NLDR) techniques
NLDR techniques designed to capture the complex and non-linear relationships present within high-dimensional data.
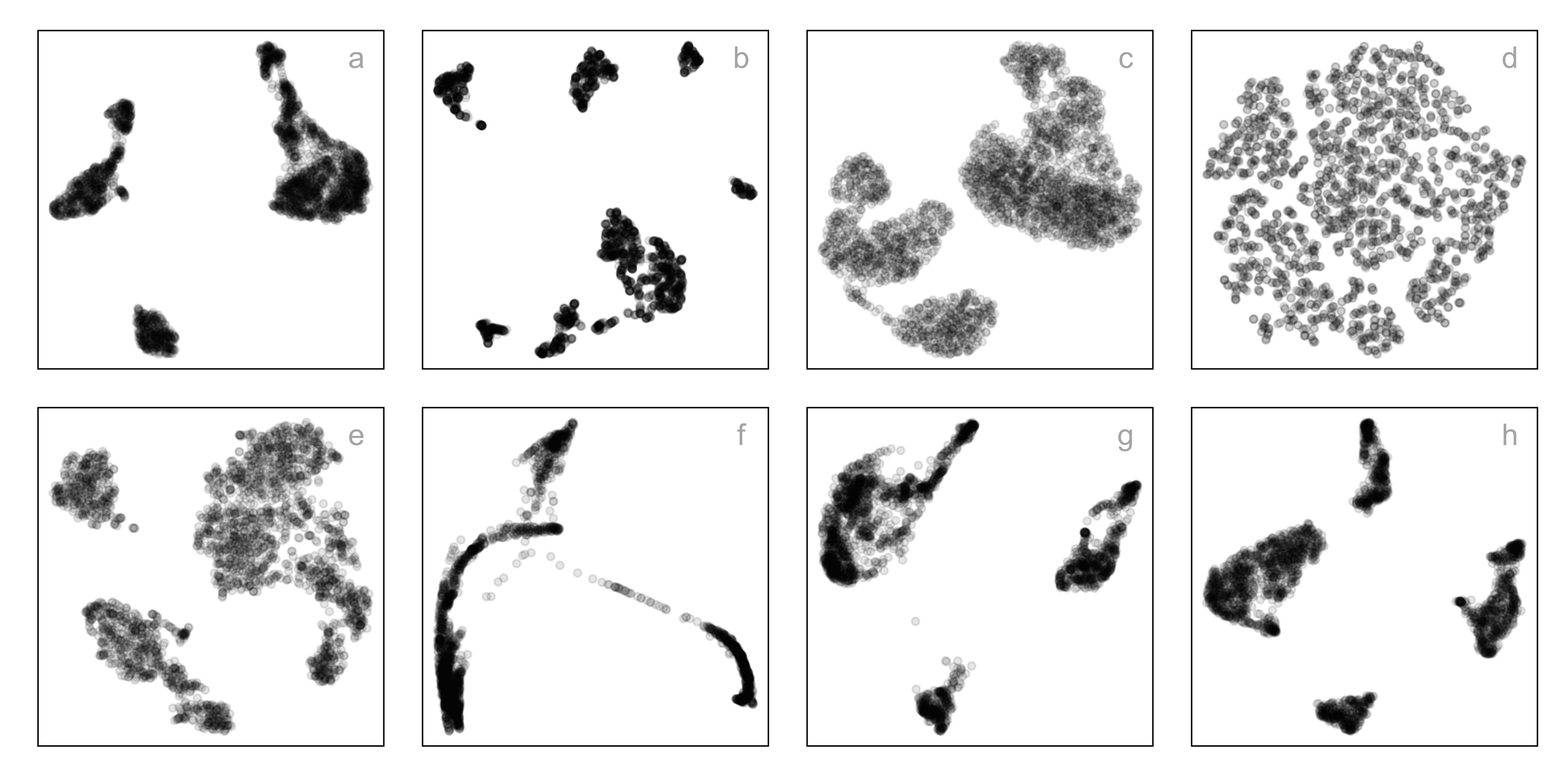
Match-a-roo (1/4)
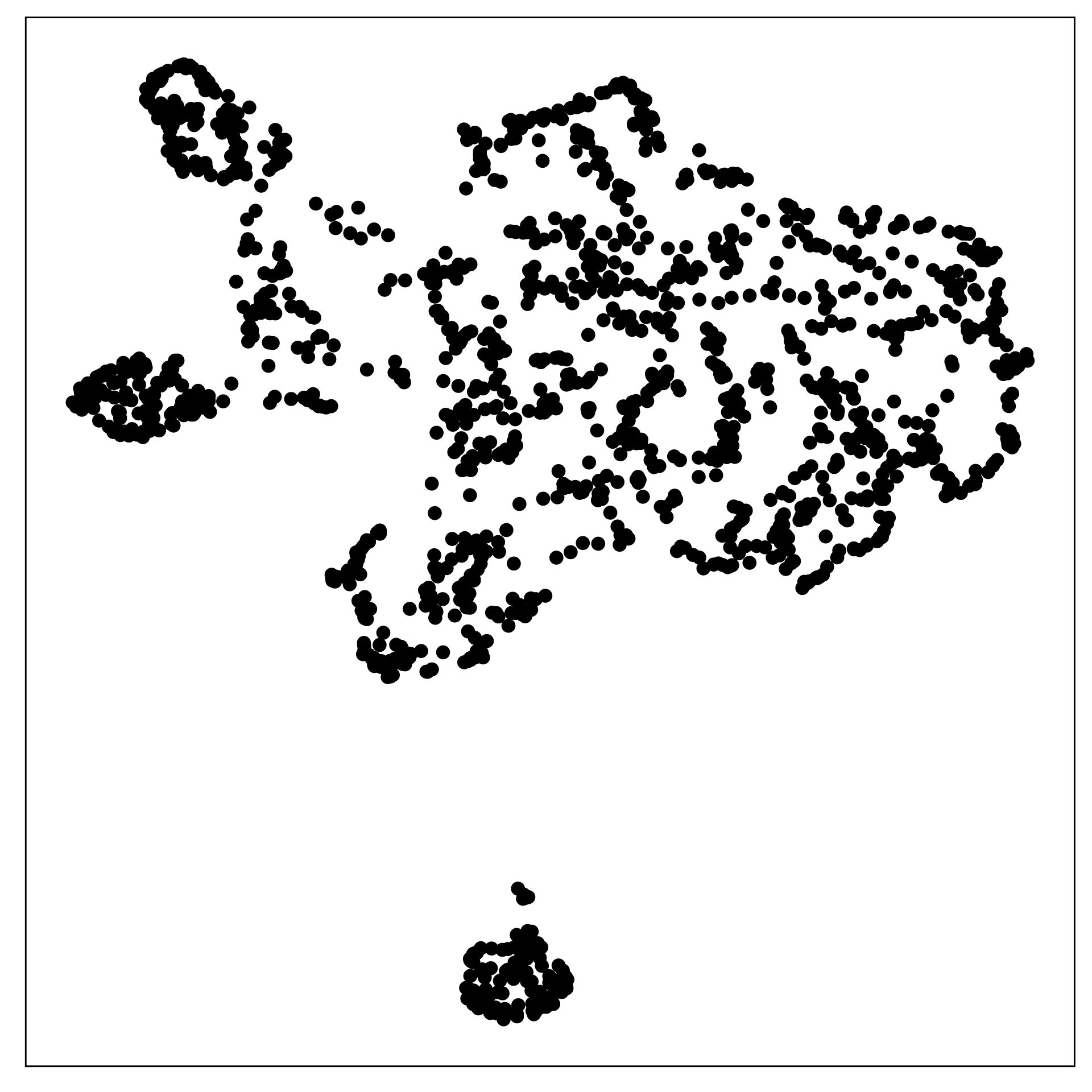
The data shown in the two displays is the
- SAME
- DIFFERENT
Match-a-roo (2/4)

The data shown in the two displays is the
- SAME
- DIFFERENT
Match-a-roo (3/4)
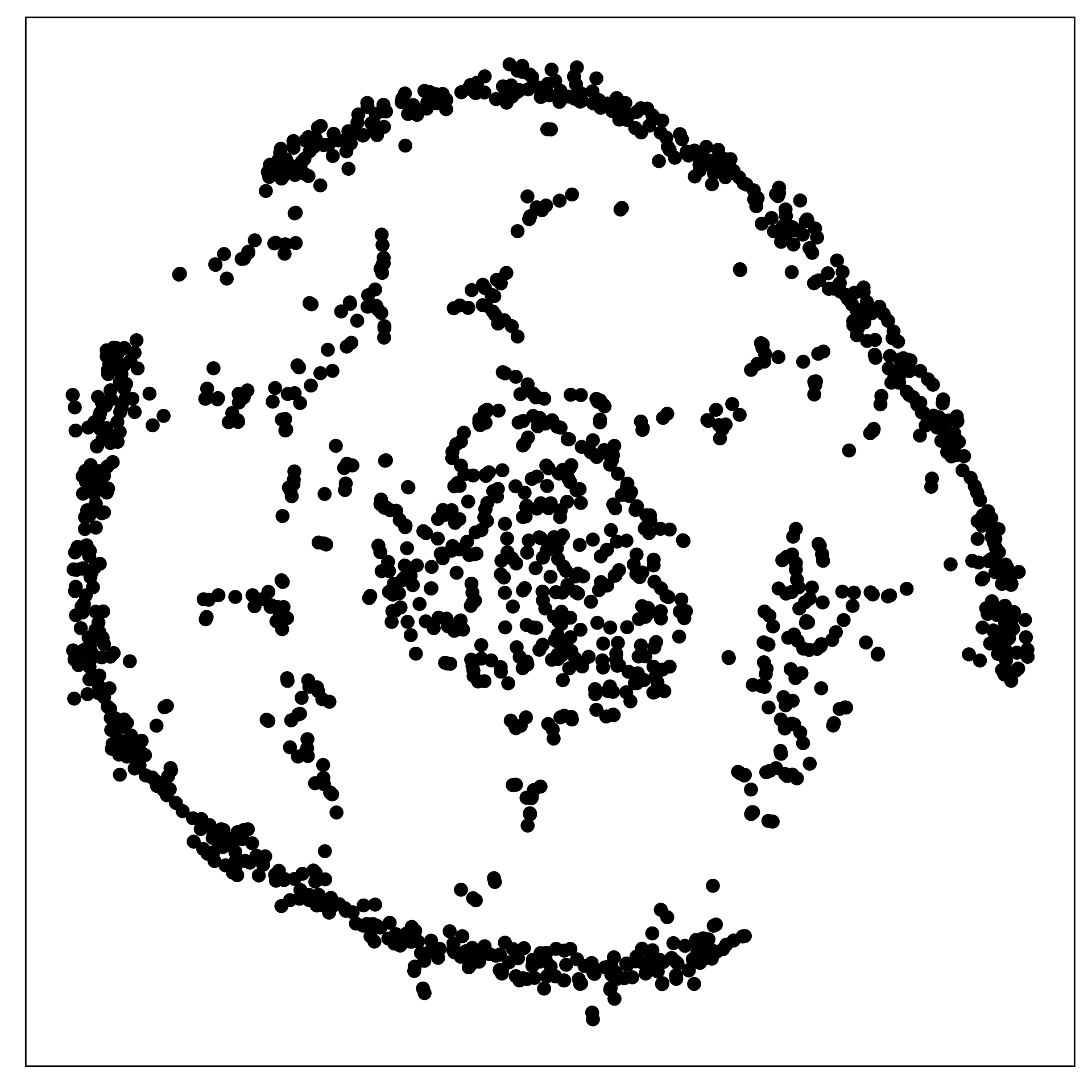
The data shown in the two displays is the
- SAME
- DIFFERENT
Match-a-roo (4/4)
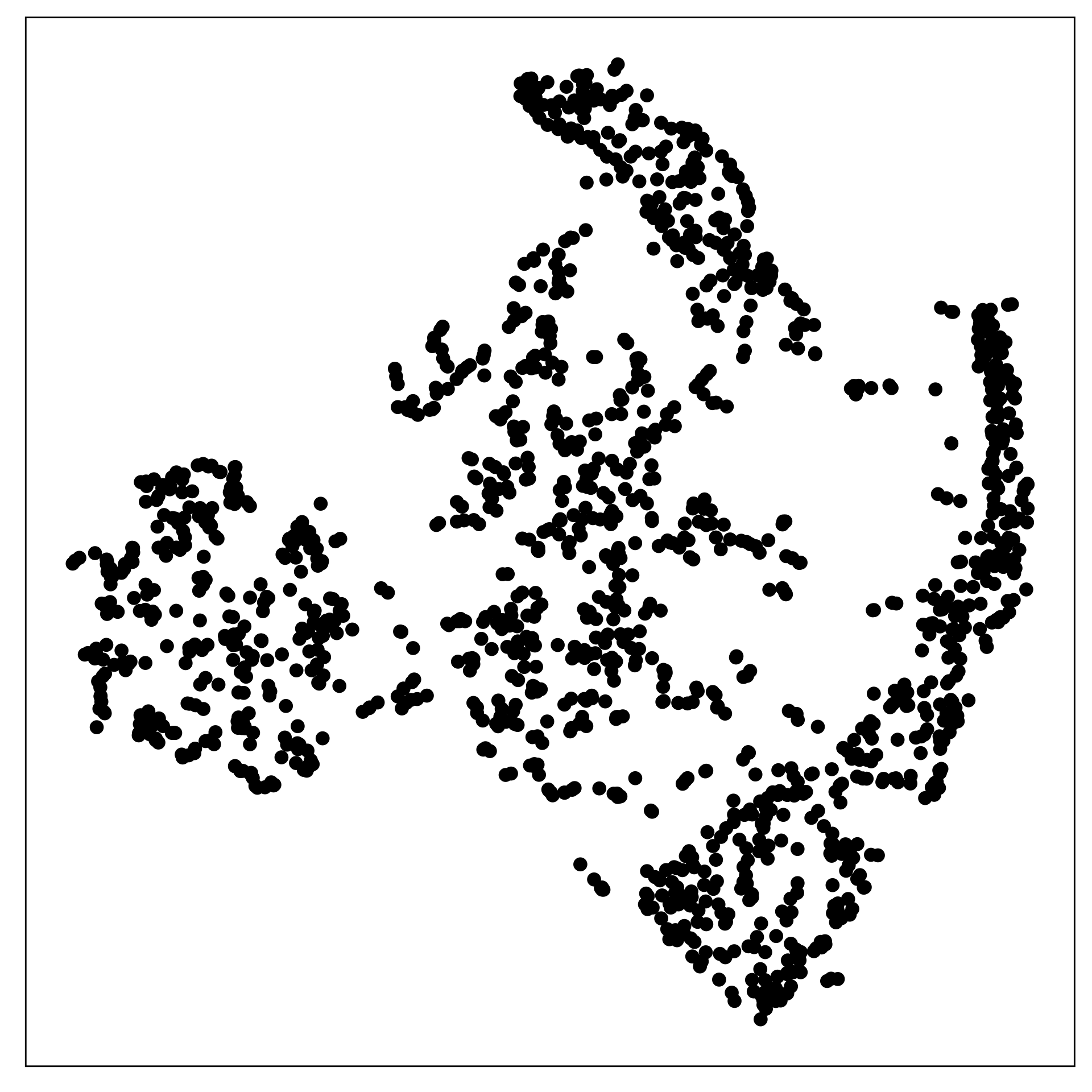
The data shown in the two displays is the
- SAME
- DIFFERENT
Motivation
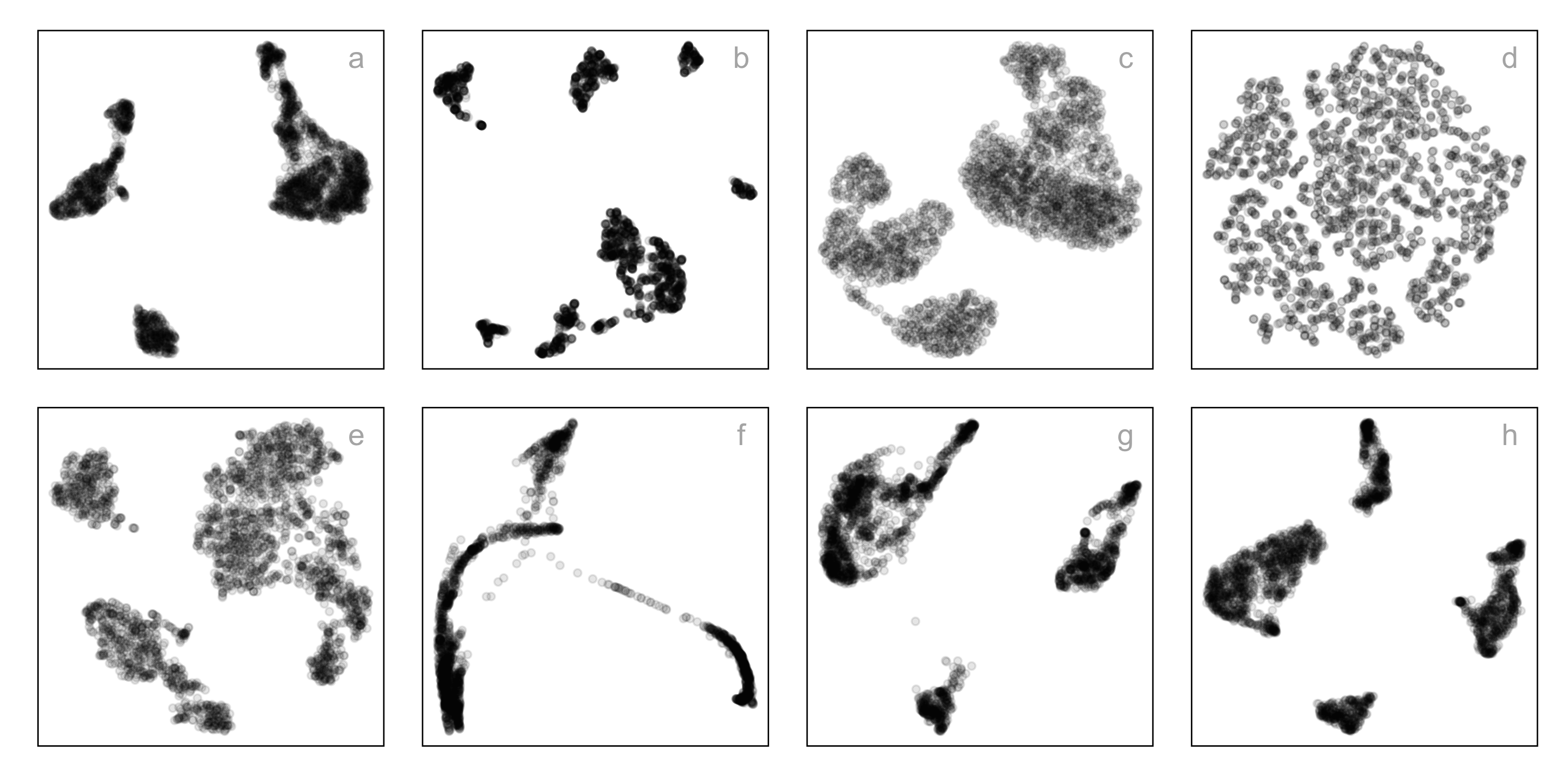
Single-cell gene expression: same data, different NLDR + hyper-parameters
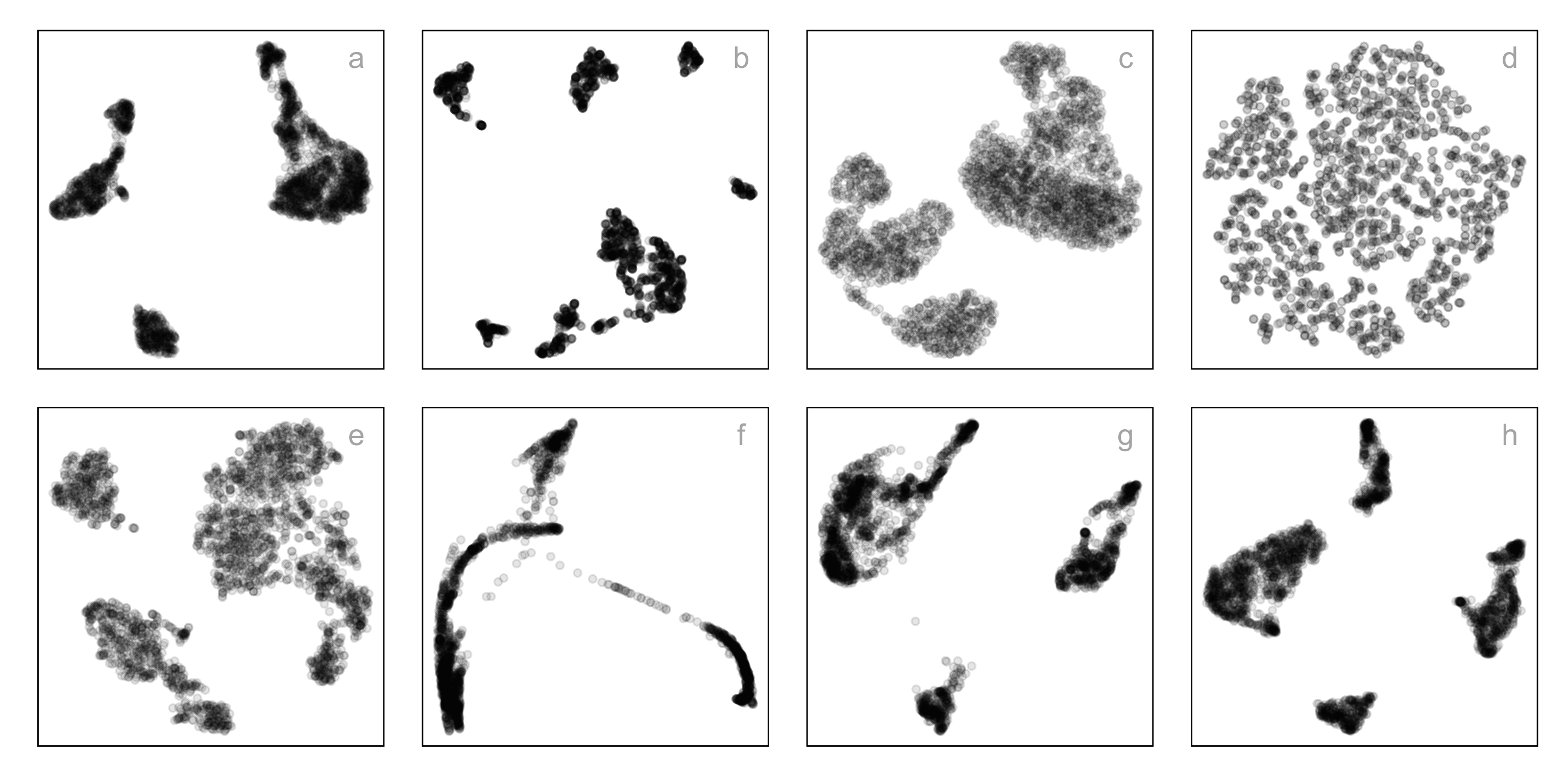
How do you decide which is the most reasonable representation?
This is the published figure.
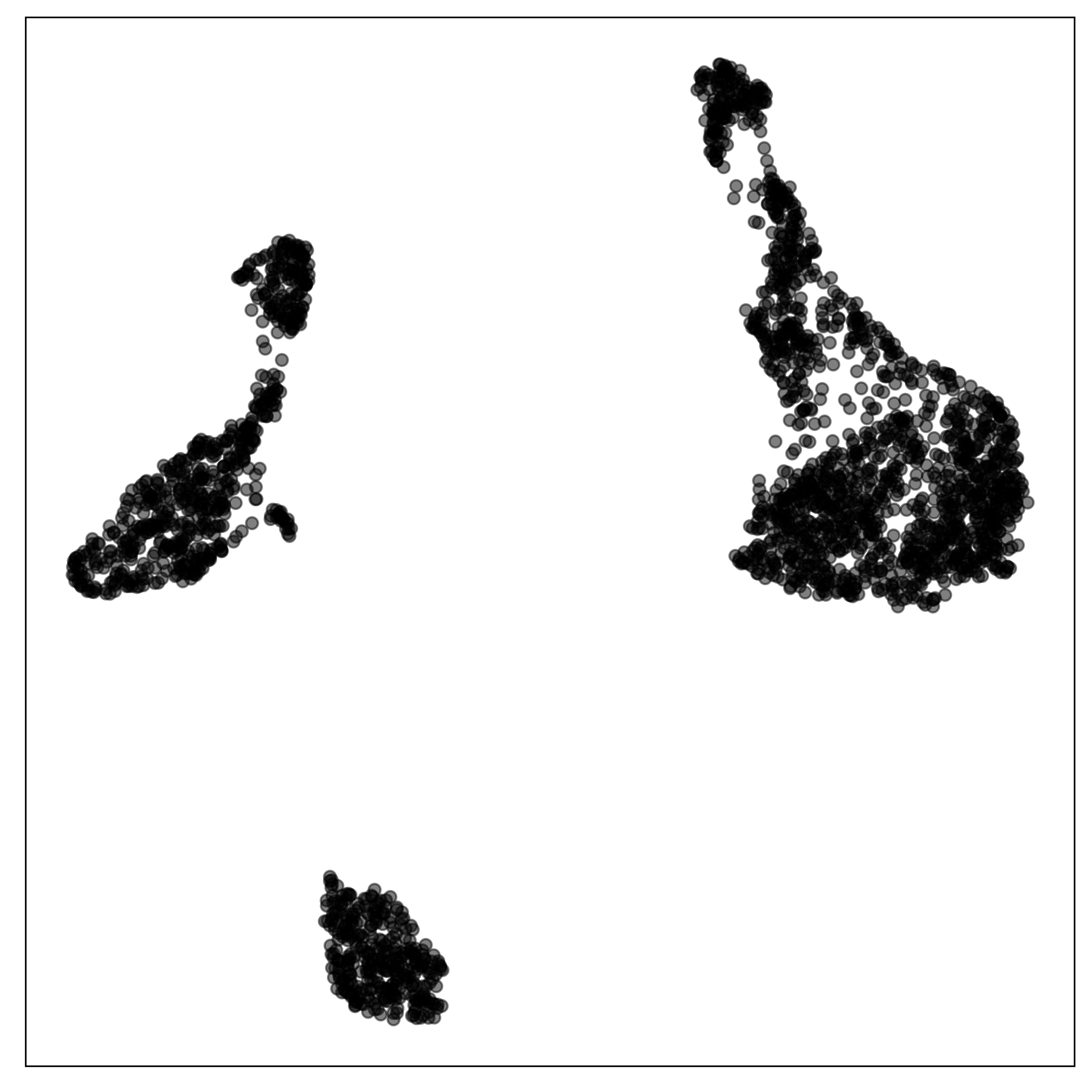

Here is the 9D data viewed using a grand tour, linear projections into 2D.
Show “model-in-the-data-space”
data-in-the-model-space
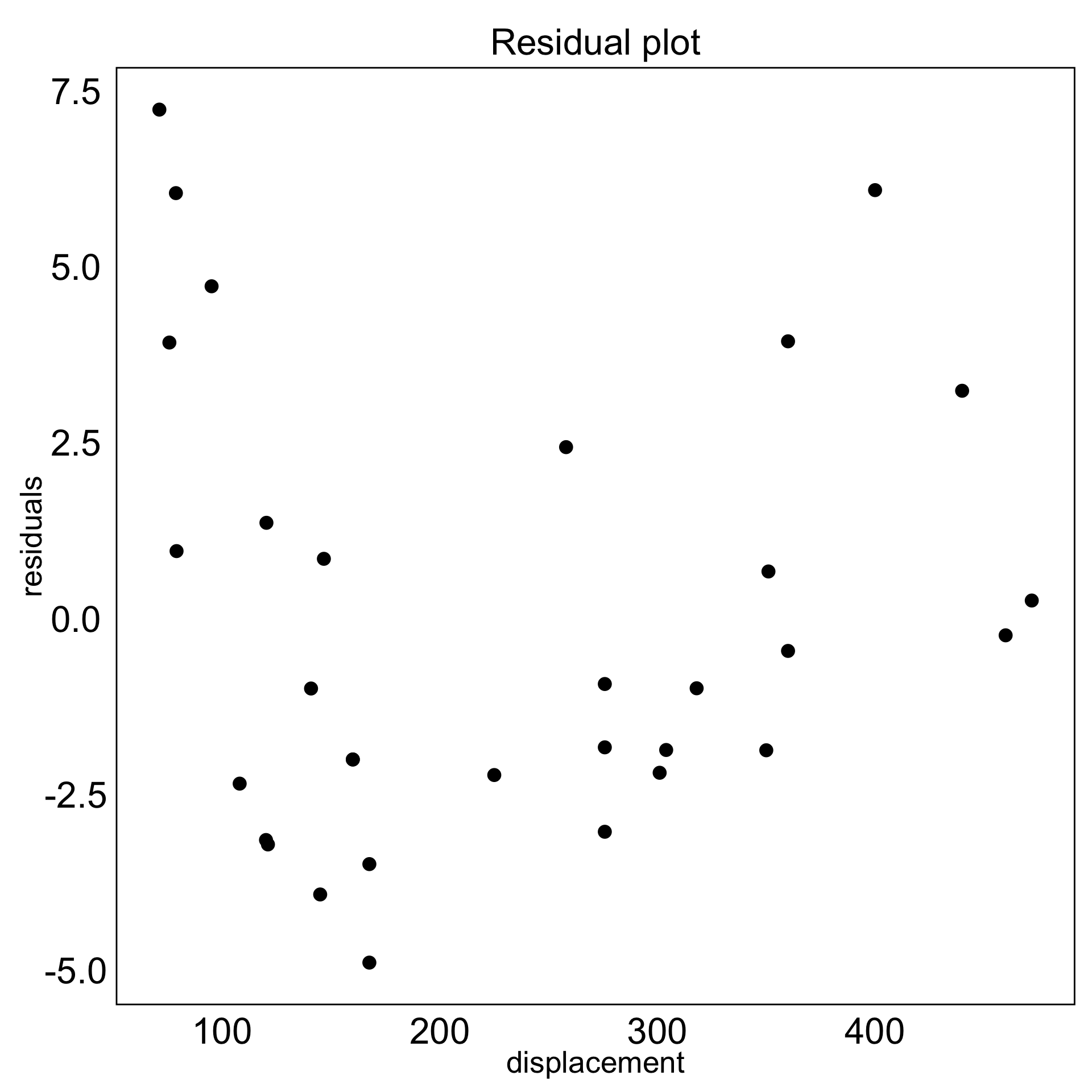
model-in-the-data-space
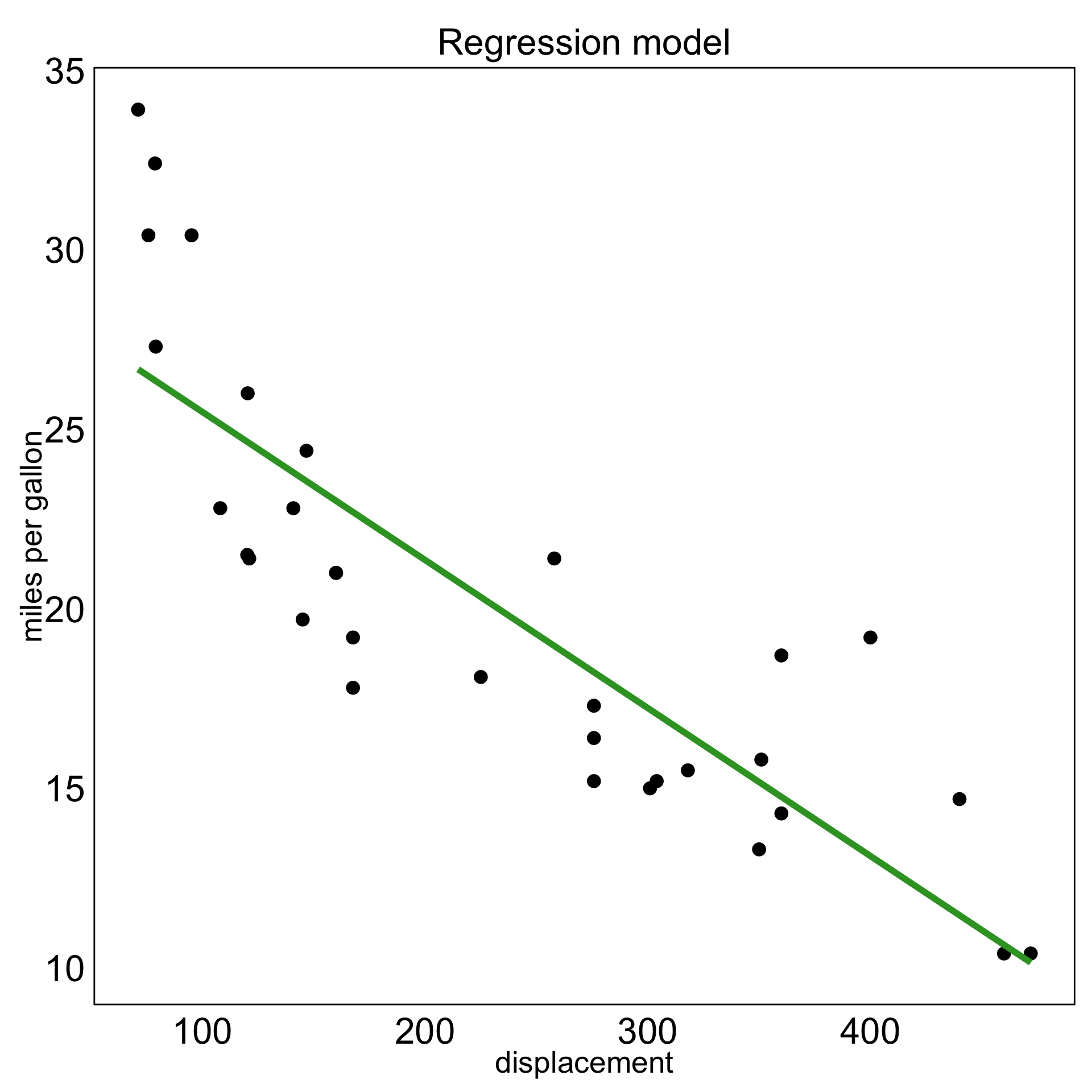
S-curve in 7D
\(\theta \sim U(-3\pi/2, 3\pi/2)\)
\(X_1 = \sin(\theta)\)
\(X_2 \sim U(0, 2)\)
\(X_3 = \text{sign}(\theta) \times (\cos(\theta) - 1)\)
True model: \(T=(X_1, X_2, X_3)\)
\(X_4, X_5, X_6, X_7\) are additional noise dimensions
data-in-the-model-space
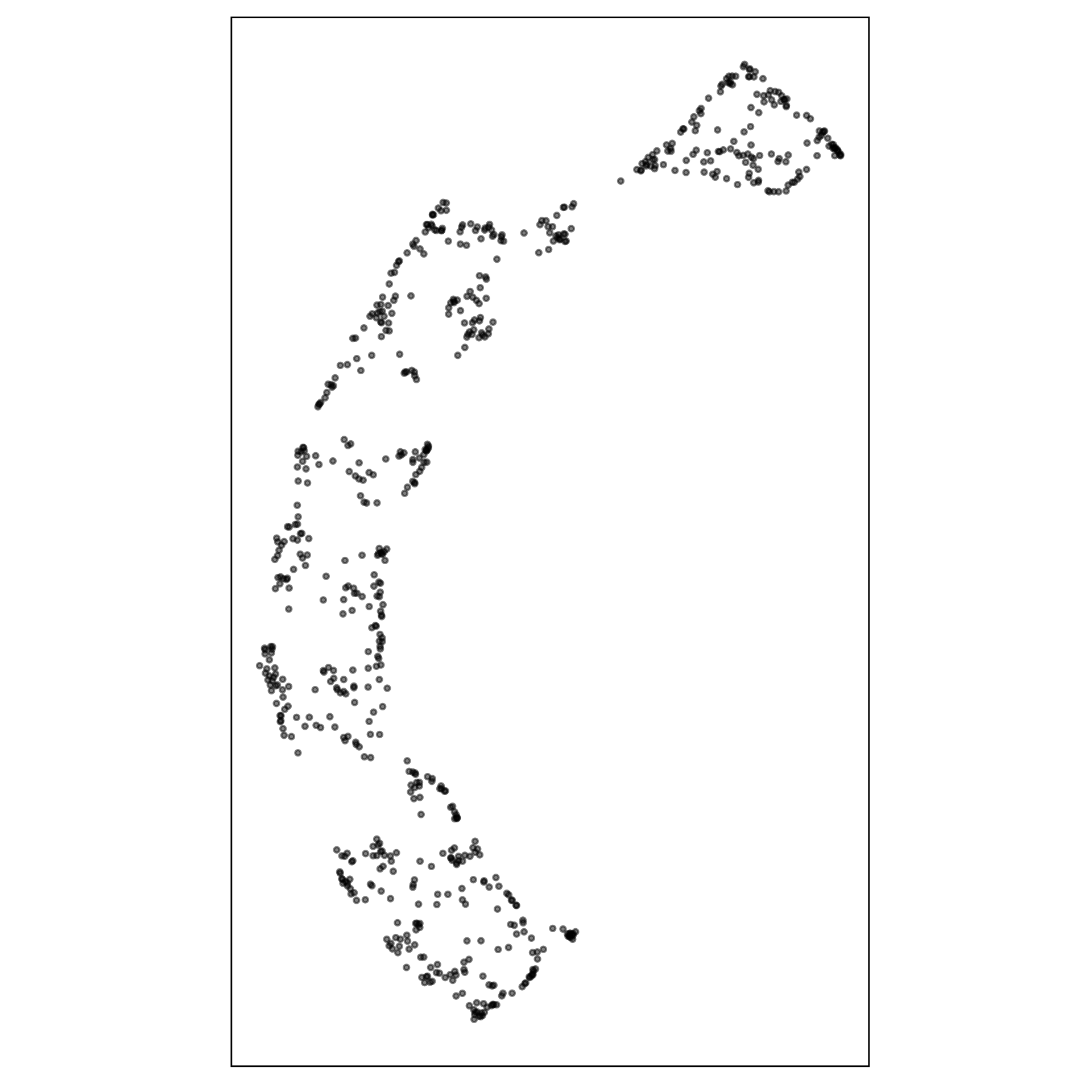
What is the model?
data-in-the-model-space
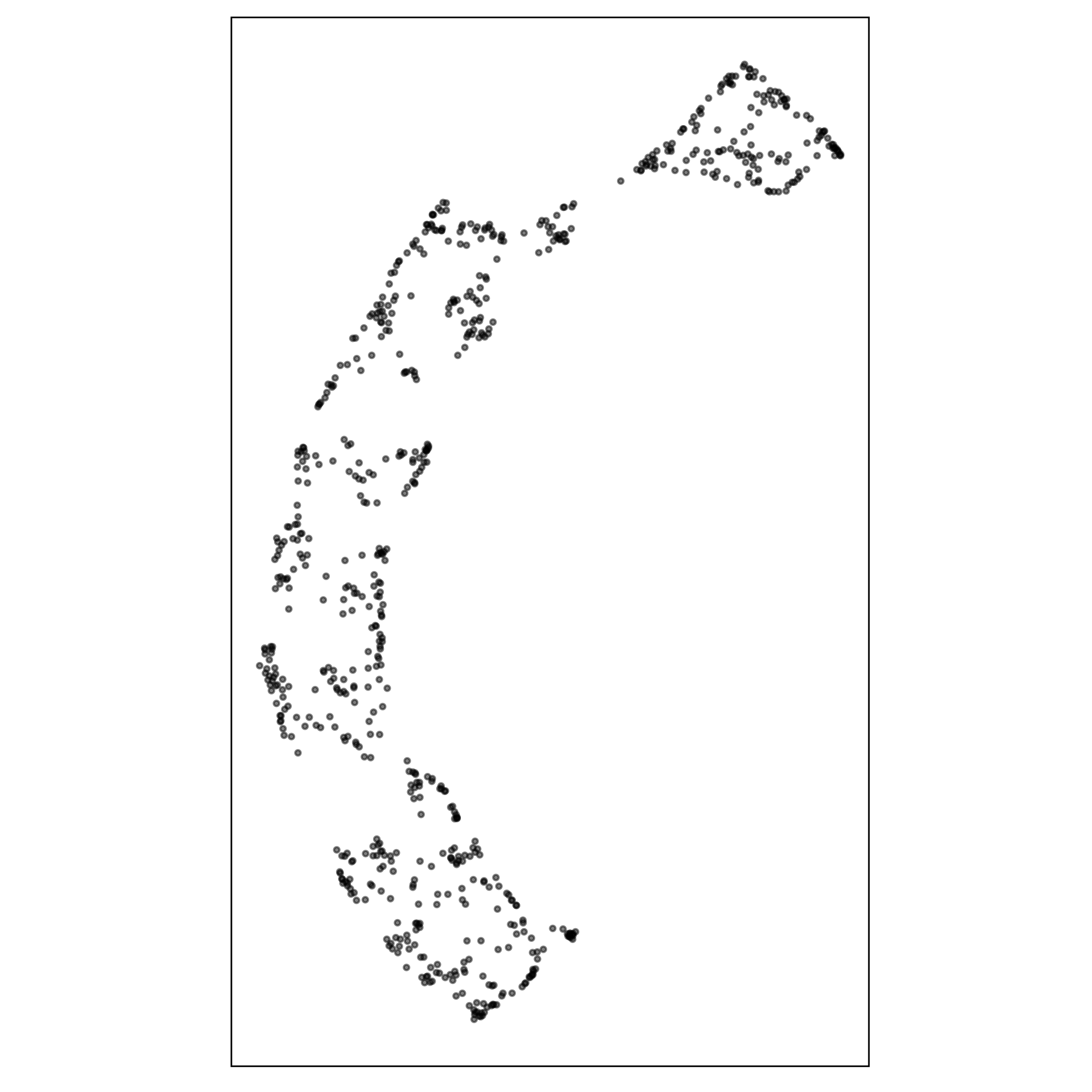
model-in-the-data-space
Overview of method
1. Construct the \(2\text{-}D\) model
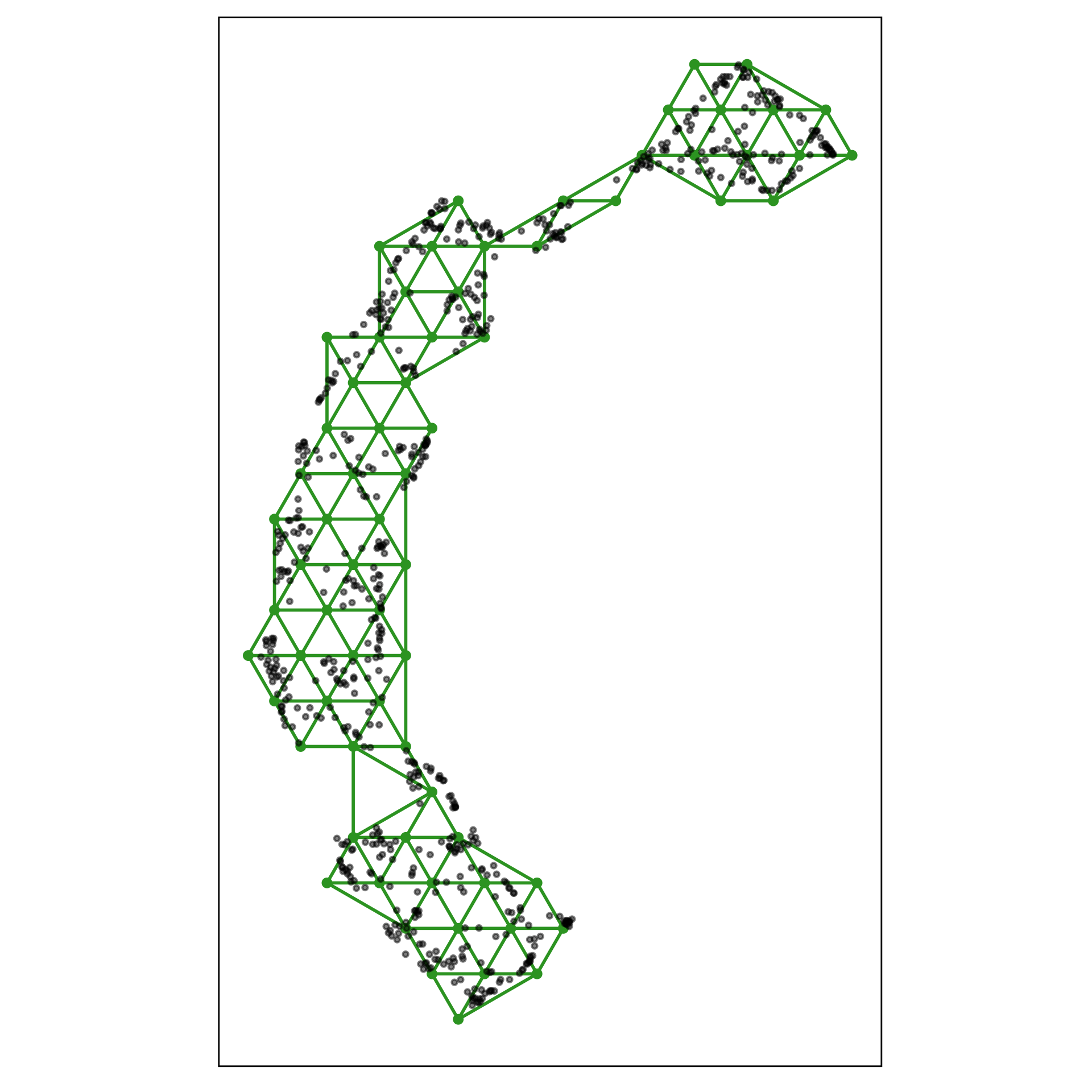
2. Lift the model into high-dimensions
Steps of the algorithm
1. Construct the \(2\text{-}D\) model
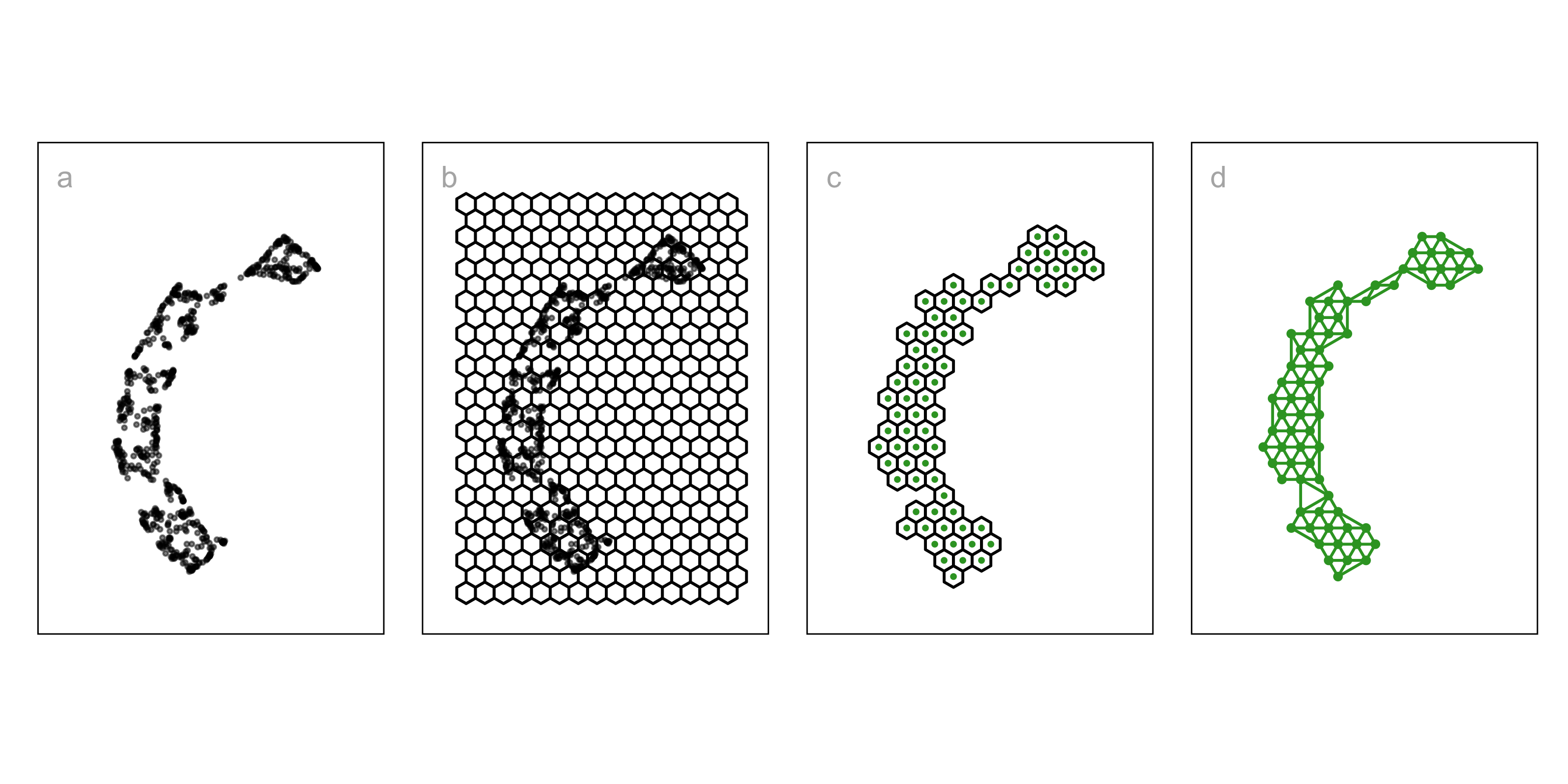
- NLDR layout, b. hex bin , c. bin centers, d. triangulation wire frame.
Steps of the algorithm
2. Lift the model into high-dimensions
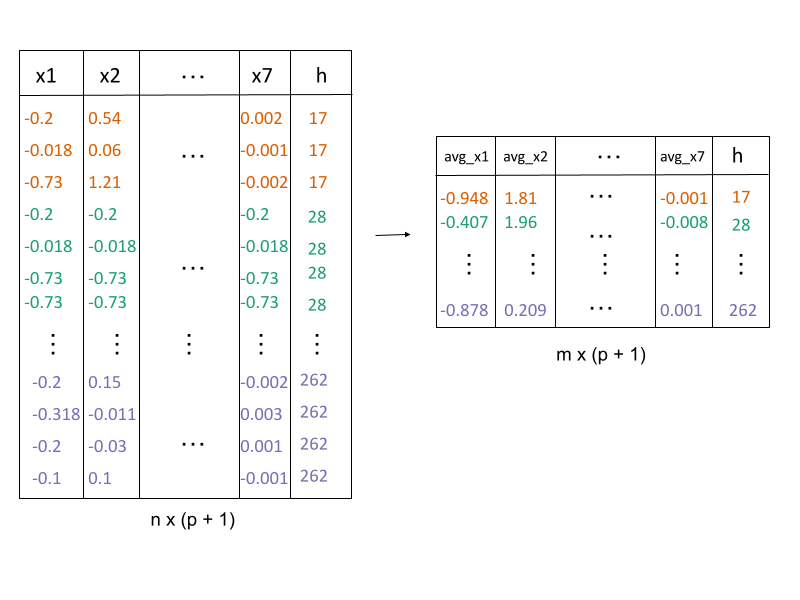
Factors for fitting and measuring fit
- NLDR layout, different methods and different hyper-parameters
- Number of bins
- Bin start position
- Low density removal
- Long edge removal
- MSE in high-dimensions: mean sum of squared differences between observed and fitted values
\[\frac{1}{n}\sum_{h = 1}^{b}\sum_{i = 1}^{n_h}\sum_{j = 1}^{p} ({x}_{hij} - C^{(p)}_{hj})^2\] \(n =\) the number of observations,
\(b =\) the number of bins,
\(n_h =\) the number of observations in \(h^{th}\) bin,
\(p =\) the number of variables,
\({x}_{hij} =\) the \(j^{th}\) dimensional data of \(i^{th}\) observation in \(h^{th}\) hexagon.
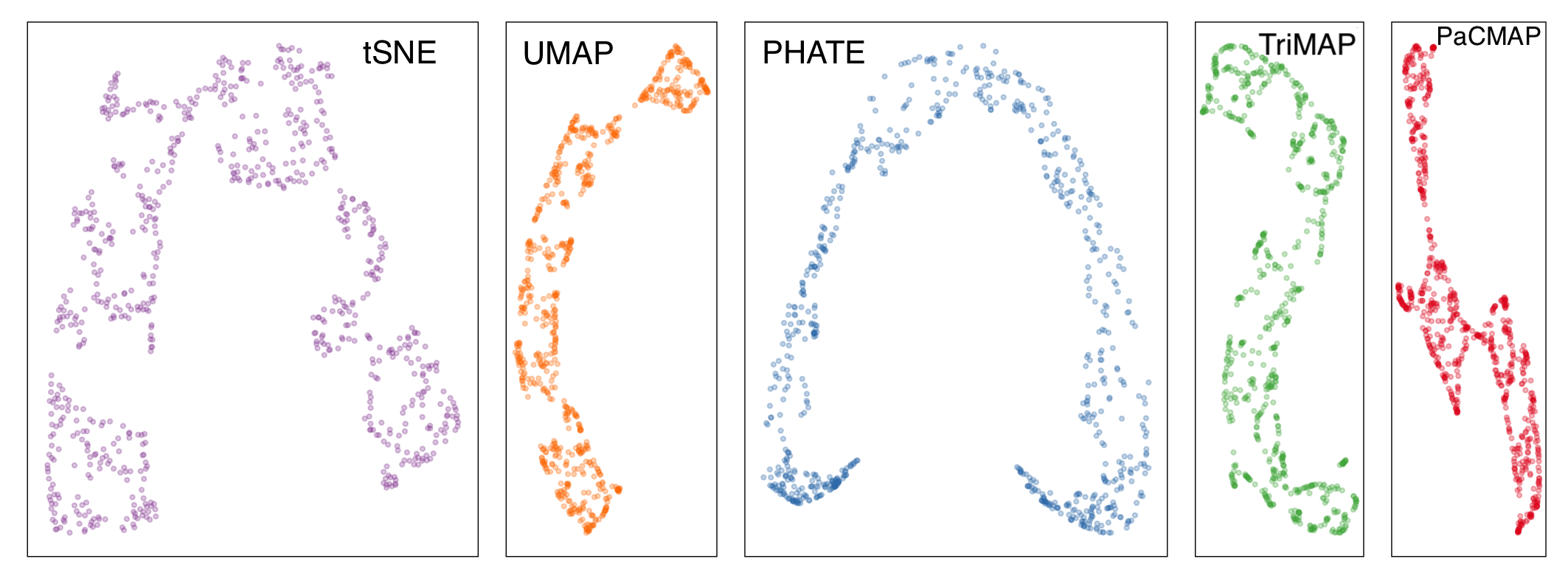
Candidates for NLDR layout
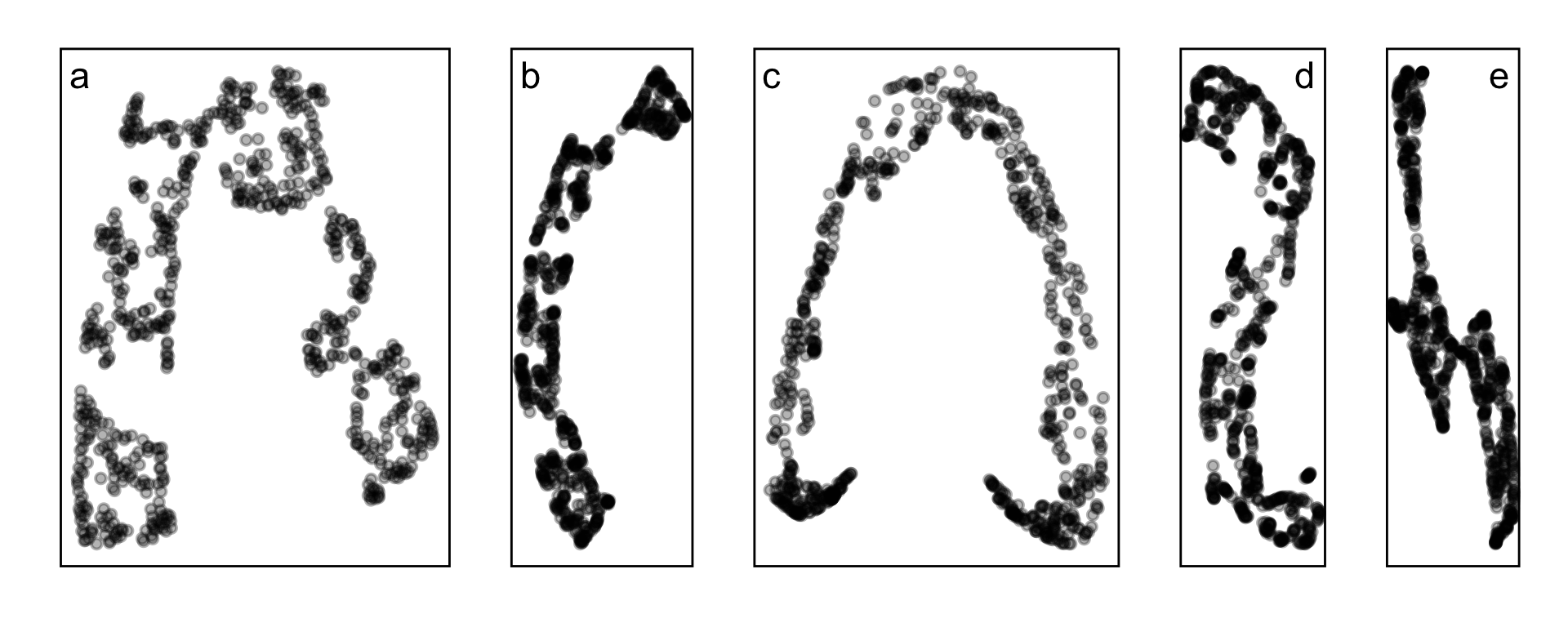
- tSNE, b. UMAP, c. PHATE, d. TriMAP, e. PaCMAP
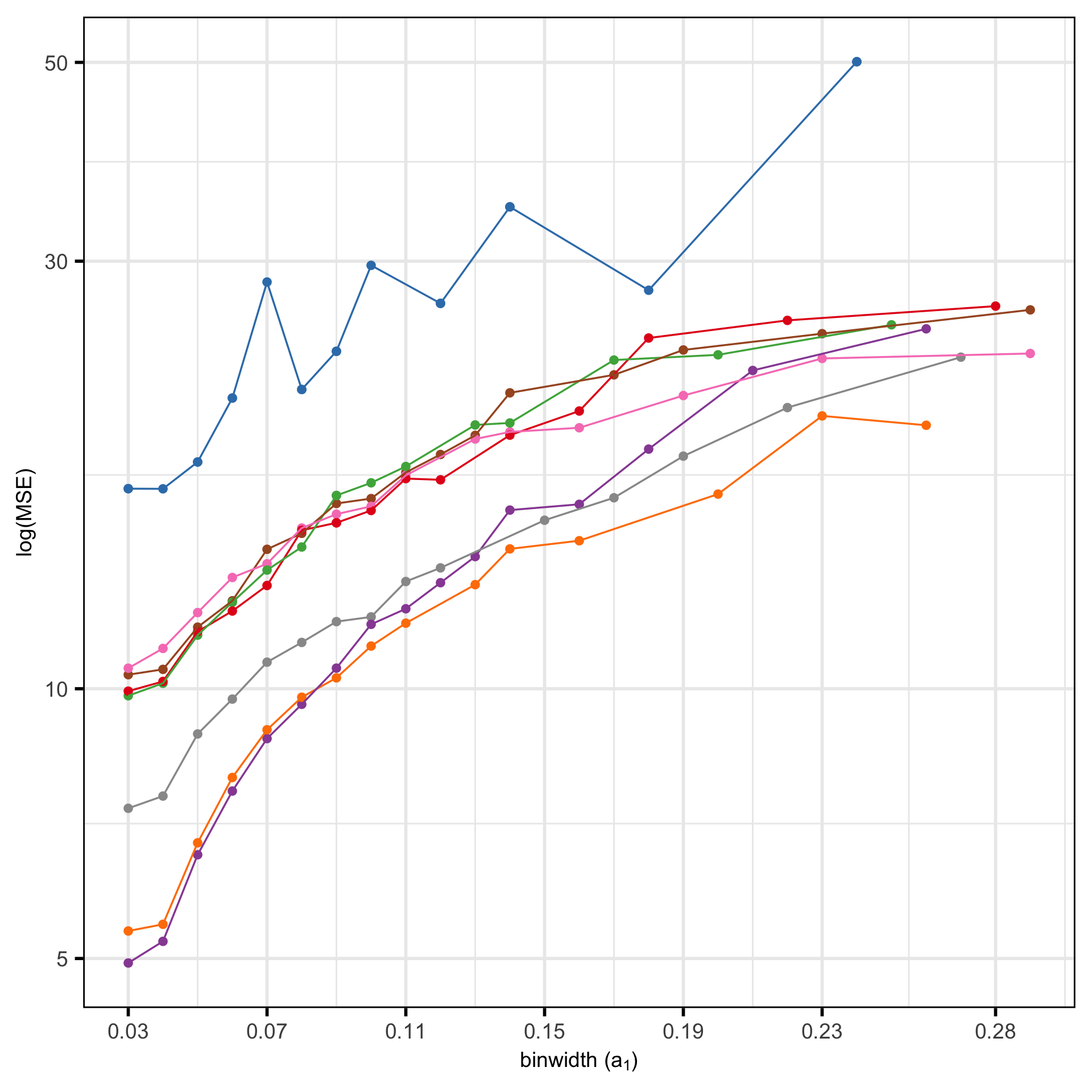
MSE of candidates
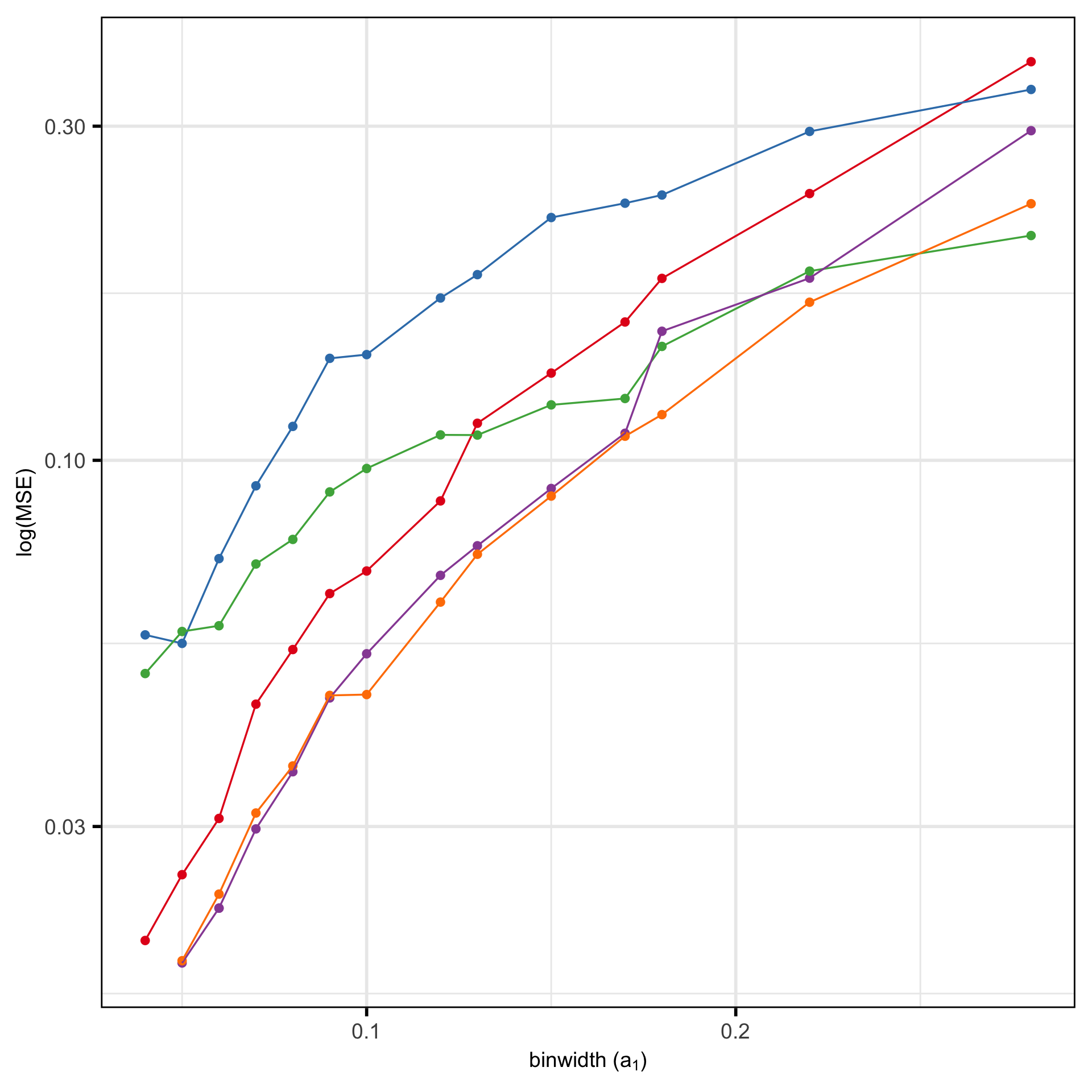
- PHATE not competitive
- Not much difference between any other method based on Error
- No elbow, just gradual decrease as number of (non-empty) bins increase
Chosen fit for S-curve
tSNE with perplexity: 27
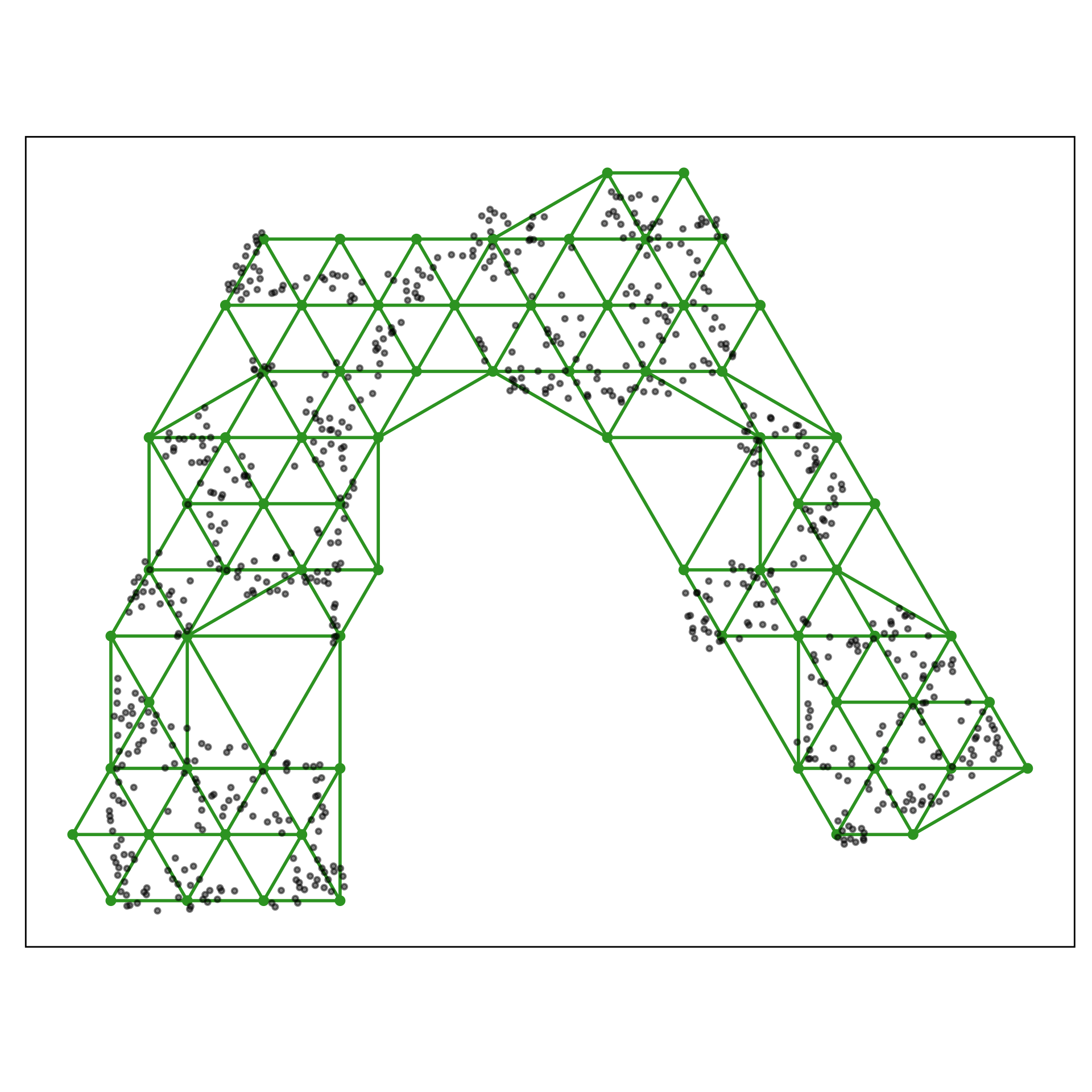
Fills out the width of the S
Pretty good! Can you see the twist??
PBMC data set
MSE of candidates
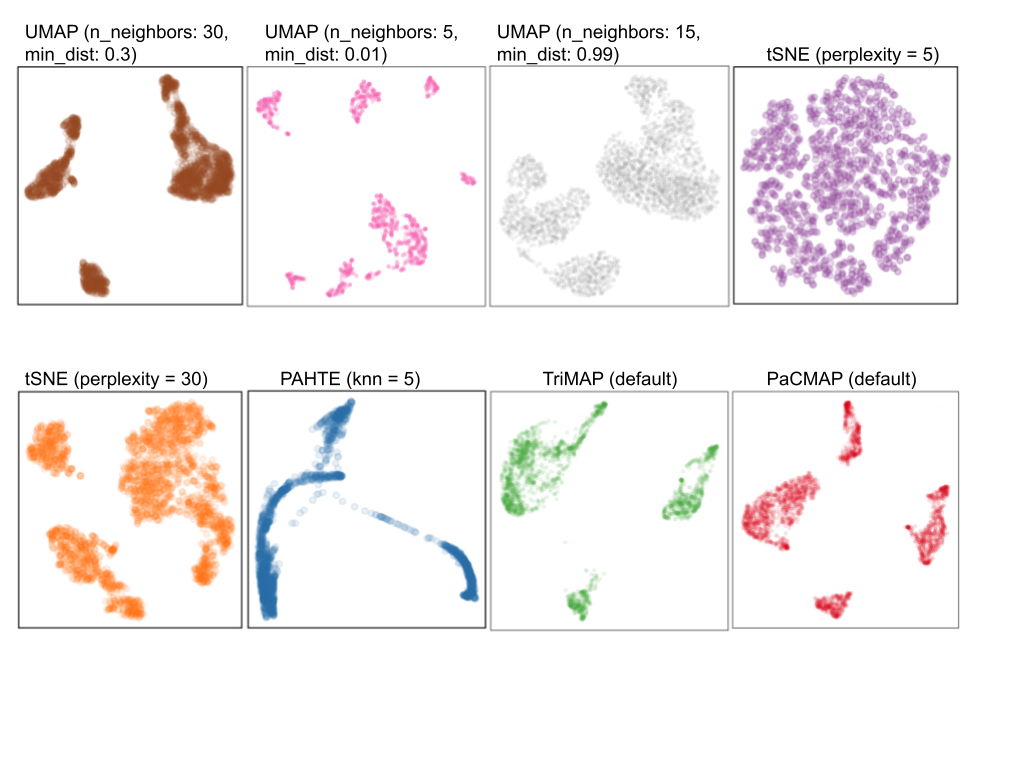
Chosen fit for PBMC data set
tSNE with perplexity: 30
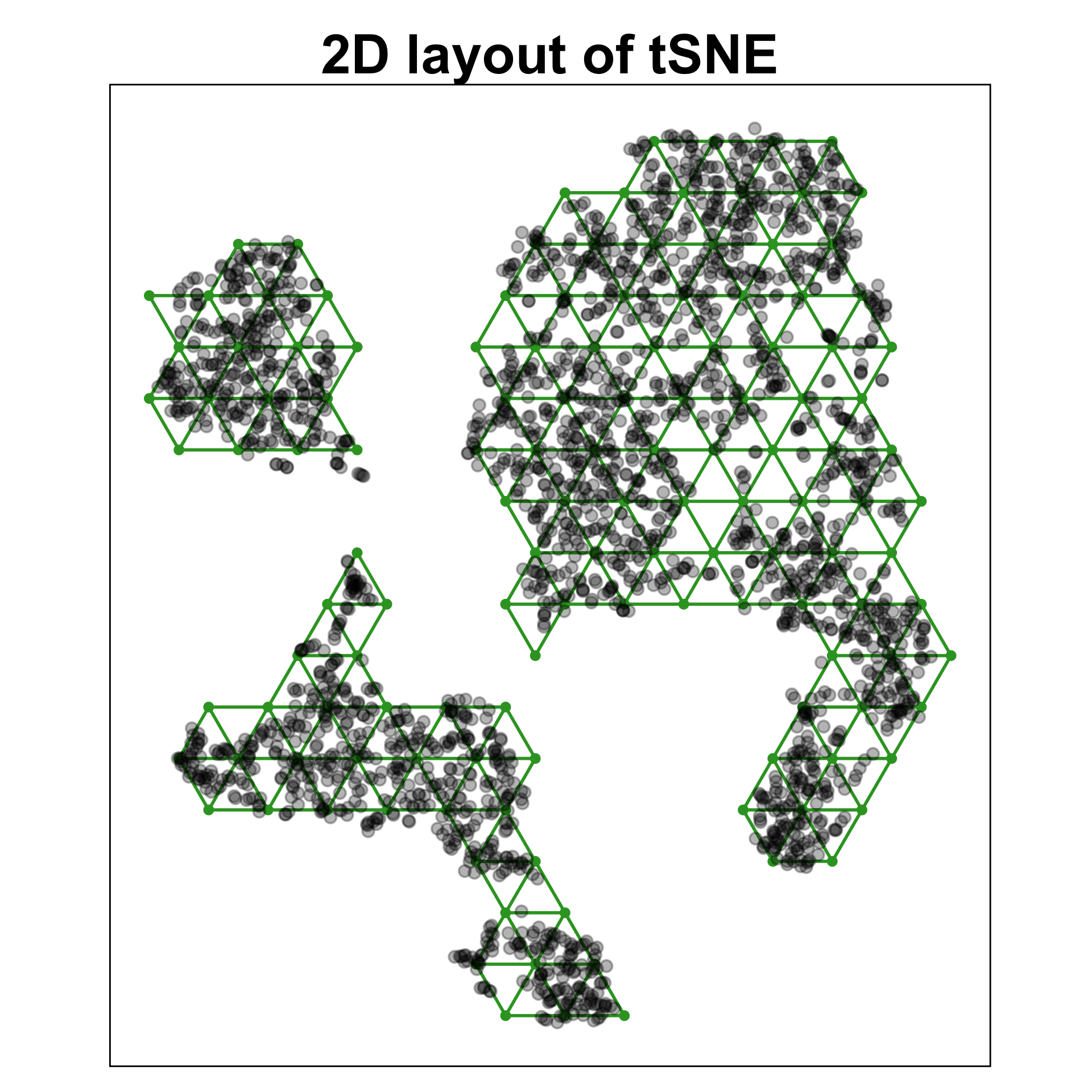
Clusters with small separations, non-linear clusters
Densed points, filled out clusters
Five Gaussian clusters in 4D
tSNE
PaCMAP
Linked plots
- Assess the model fits the points everywhere, better in some places, simply mismatches the pattern


quollr
questioning how a high-dimensional object looks in low-dimensions using r
Summary
Note
- Provided a method to create a model from a NLDR layout that can be displayed with the data to assess the fit.
- Make it easier for researchers to make better decisions on which NLDR layout is best for their work.

Jayani P.G. Lakshika 
Collaborators: Prof. Dianne Cook, Dr. Paul Harrison, Dr. Michael Lydeamore, Dr. Thiyanga S. Talagala